Science AI
Inspired by the ability of AI to help tackle the grand challenges in science, our work involves pursuing breakthrough discoveries and developing new tools to accelerate the progress of science. Together with the scientific community, our aim is to enable scientific innovation for a better world.
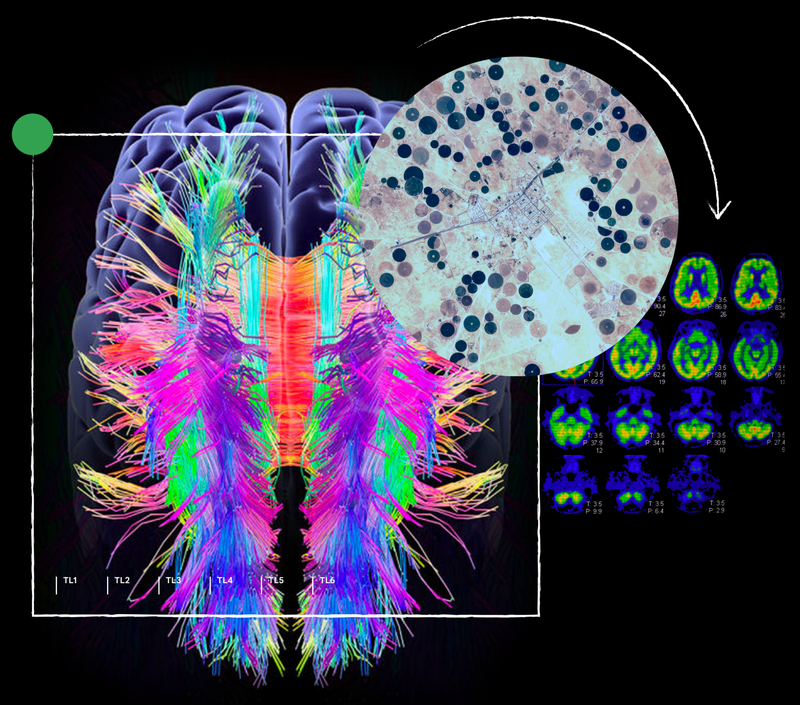
Inspired by the ability of AI to help tackle the grand challenges in science, our work involves pursuing breakthrough discoveries and developing new tools to accelerate the progress of science. Together with the scientific community, our aim is to enable scientific innovation for a better world.
-
Collaborative and open science
Our history of scientific contributions is rooted in highly collaborative research that promotes transparent, accessible and rigorous results. We develop open source software, release open datasets and partner with labs globally to co-create breakthroughs.
-
Tools to accelerate scientific discovery
We build AI-powered tools to empower the global scientific community to quickly move from idea to discovery. These experimental tools, including Empirical Research Assistance and AI co-scientist, help scientists tackle complex questions once thought impossible, and solve the world’s most daunting challenges at unprecedented speed.
Decoding biological complexity
We use AI to tackle the biggest unsolved questions in biology, from understanding the genome to solving the mysteries of the brain. These insights address core questions that benefit biomedical researchers, and ultimately millions of patients affected by rare genetic and neurological conditions.
-
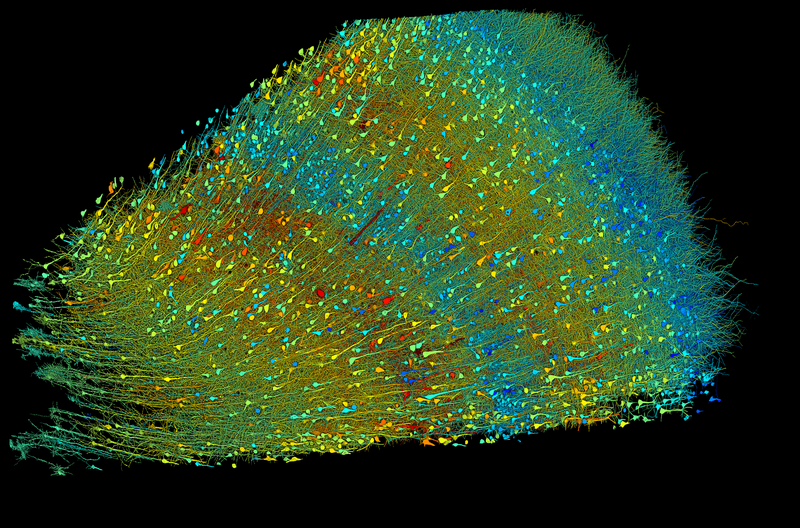 Using AI to map and understand the brain
Using AI to map and understand the brainWe are driving progress toward precisely mapping the connections between every cell in the brain. Through an evolving set of AI-based tools for neural reconstruction, analysis, and visualization, we help scientists unlock a deeper understanding of how we think, learn and experience the world.
-
 Genomics: Using AI to interpret the genome
Genomics: Using AI to interpret the genomeWe pioneered the use of deep learning in genomics, and now partner with labs globally to decode the DNA of humans, animals and plants. Our research enables discoveries and advances healthcare while also providing critical tools to preserve the world’s biodiversity.
Modeling the earth & environment
We use AI to improve humanity’s fundamental understanding of Earth’s complex systems: water, land, life and sky. By understanding our planet, we enable communities, researchers and governments to make more informed decisions that foster resilience and achieve a safer, healthier and more sustainable future for all.
-
 Flood forecasting
Flood forecastingOur flood forecasting models use AI to help close the global warning gap, providing reliable flood predictions even in regions that previously lacked data.
-
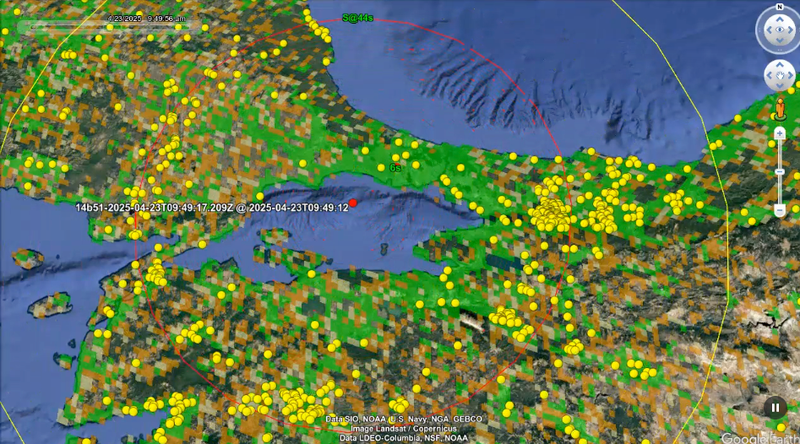 Earthquake detection
Earthquake detectionWe leverage measurements from a global network of Android smartphones to detect earthquakes and deliver early warnings to people in affected areas.
-
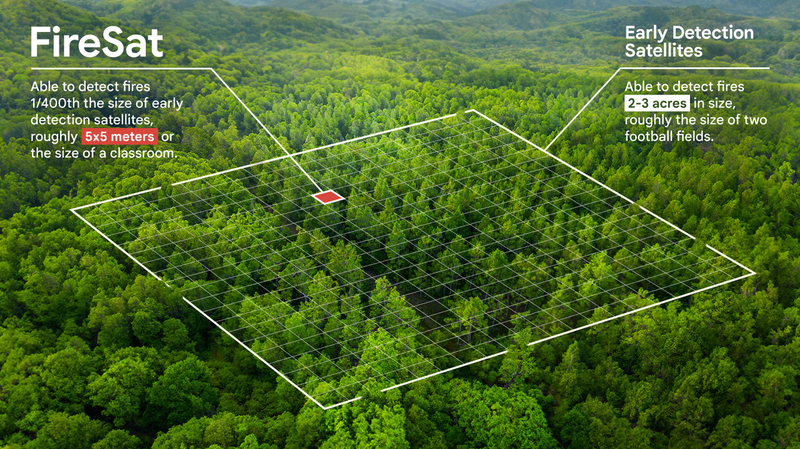 Wildfire research
Wildfire researchWe use satellite imagery and AI to detect and track the boundaries of wildfires, simulate wildfire evolution, and provide near real-time alerts to help affected communities and responders act fast.
-
 Species mapping
Species mappingAs Earth’s biodiversity faces a crisis, we are using AI to map plant, animal and fungi distributions at unprecedented scale. These maps help identify where species live, providing critical data for conservation efforts.
-
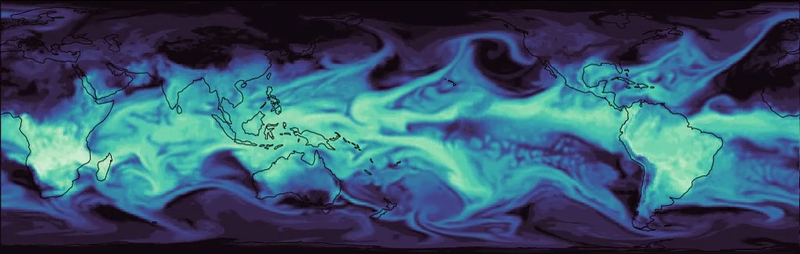 Atmospheric modeling
Atmospheric modelingOur NeuralGCM model combines the best of physics and AI to rapidly, efficiently, and accurately simulate Earth’s atmosphere. This hybrid approach enables researchers to run climate simulations more efficiently than ever before.
-
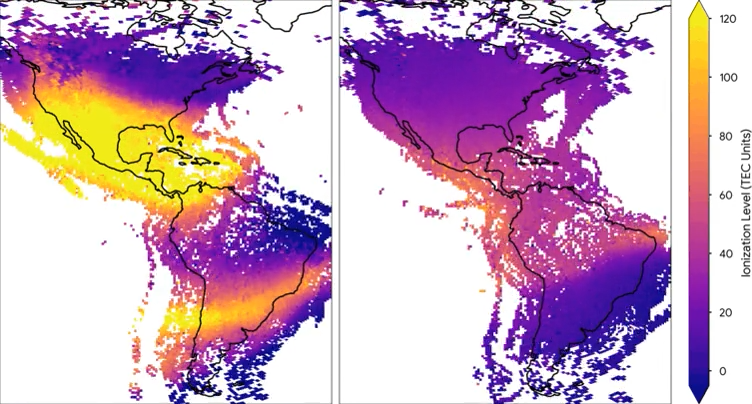 Mapping space weather
Mapping space weatherWe used aggregated measurements from millions of Android phones to measure the ionosphere and space weather in Earth’s upper atmosphere, doubling global coverage compared to traditional methods.
Recent featured publications
Explore and engage with our research via curated NotebookLMs
Our Broader Mission
-
 Support for scientists
Support for scientistsGoogle.org brings the best of Google to organizations around the globe by providing funding, programs, and technical expertise to accelerate their missions.
-
 Core AI principles
Core AI principlesOur discoveries exemplify Google’s core principles for AI-driven innovation: bold, responsible and collaborative.



