Computer-aided diagnosis for lung cancer screening
March 20, 2024
Posted by Atilla Kiraly, Software Engineer, and Rory Pilgrim, Product Manager, Google Research
Lung cancer is the leading cause of cancer-related deaths globally with 1.8 million deaths reported in 2020. Late diagnosis dramatically reduces the chances of survival. Lung cancer screening via computed tomography (CT), which provides a detailed 3D image of the lungs, has been shown to reduce mortality in high-risk populations by at least 20% by detecting potential signs of cancers earlier. In the US, screening involves annual scans, with some countries or cases recommending more or less frequent scans.
The United States Preventive Services Task Force recently expanded lung cancer screening recommendations by roughly 80%, which is expected to increase screening access for women and racial and ethnic minority groups. However, false positives (i.e., incorrectly reporting a potential cancer in a cancer-free patient) can cause anxiety and lead to unnecessary procedures for patients while increasing costs for the healthcare system. Moreover, efficiency in screening a large number of individuals can be challenging depending on healthcare infrastructure and radiologist availability.
At Google we have previously developed machine learning (ML) models for lung cancer detection, and have evaluated their ability to automatically detect and classify regions that show signs of potential cancer. Performance has been shown to be comparable to that of specialists in detecting possible cancer. While they have achieved high performance, effectively communicating findings in realistic environments is necessary to realize their full potential.
To that end, in “Assistive AI in Lung Cancer Screening: A Retrospective Multinational Study in the US and Japan”, published in Radiology AI, we investigate how ML models can effectively communicate findings to radiologists. We also introduce a generalizable user-centric interface to help radiologists leverage such models for lung cancer screening. The system takes CT imaging as input and outputs a cancer suspicion rating using four categories (no suspicion, probably benign, suspicious, highly suspicious) along with the corresponding regions of interest. We evaluate the system’s utility in improving clinician performance through randomized reader studies in both the US and Japan, using the local cancer scoring systems (Lung-RADSs V1.1 and Sendai Score) and image viewers that mimic realistic settings. We found that reader specificity increases with model assistance in both reader studies. To accelerate progress in conducting similar studies with ML models, we have open-sourced code to process CT images and generate images compatible with the picture archiving and communication system (PACS) used by radiologists.
Developing an interface to communicate model results
Integrating ML models into radiologist workflows involves understanding the nuances and goals of their tasks to meaningfully support them. In the case of lung cancer screening, hospitals follow various country-specific guidelines that are regularly updated. For example, in the US, Lung-RADs V1.1 assigns an alpha-numeric score to indicate the lung cancer risk and follow-up recommendations. When assessing patients, radiologists load the CT in their workstation to read the case, find lung nodules or lesions, and apply set guidelines to determine follow-up decisions.
Our first step was to improve the previously developed ML models through additional training data and architectural improvements, including self-attention. Then, instead of targeting specific guidelines, we experimented with a complementary way of communicating AI results independent of guidelines or their particular versions. Specifically, the system output offers a suspicion rating and localization (regions of interest) for the user to consider in conjunction with their own specific guidelines. The interface produces output images directly associated with the CT study, requiring no changes to the user’s workstation. The radiologist only needs to review a small set of additional images. There is no other change to their system or interaction with the system.
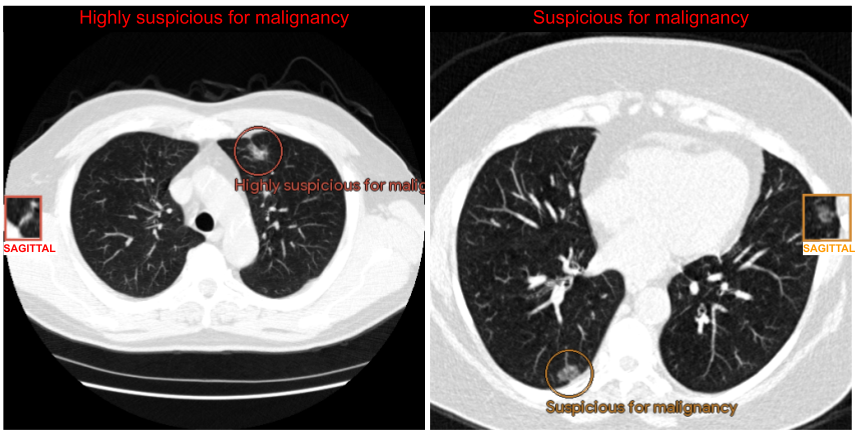
Example of the assistive lung cancer screening system outputs. Results for the radiologist’s evaluation are visualized on the location of the CT volume where the suspicious lesion is found. The overall suspicion is displayed at the top of the CT images. Circles highlight the suspicious lesions while squares show a rendering of the same lesion from a different perspective, called a sagittal view.
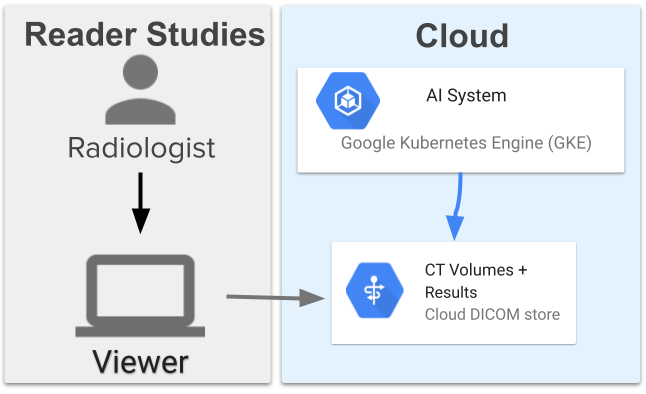
The assistive lung cancer screening system comprises 13 models and has a high-level architecture similar to the end-to-end system used in prior work. The models coordinate with each other to first segment the lungs, obtain an overall assessment, locate three suspicious regions, then use the information to assign a suspicion rating to each region. The system was deployed on Google Cloud using a Google Kubernetes Engine (GKE) that pulled the images, ran the ML models, and provided results. This allows scalability and directly connects to servers where the images are stored in DICOM stores.
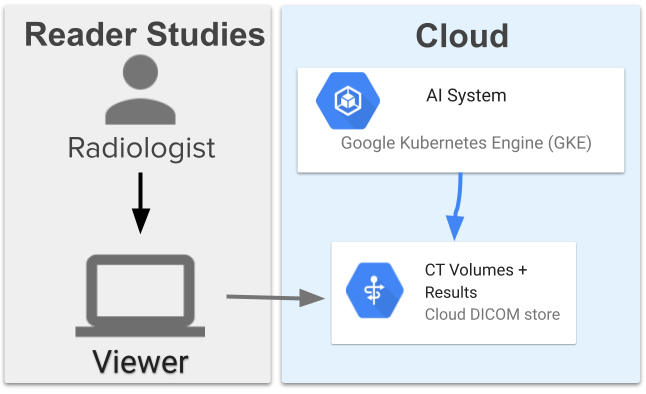
Outline of the Google Cloud deployment of the assistive lung cancer screening system and the directional calling flow for the individual components that serve the images and compute results. Images are served to the viewer and to the system using Google Cloud services. The system is run on a Google Kubernetes Engine that pulls the images, processes them, and writes them back into the DICOM store.
Reader studies
To evaluate the system’s utility in improving clinical performance, we conducted two reader studies (i.e., experiments designed to assess clinical performance comparing expert performance with and without the aid of a technology) with 12 radiologists using pre-existing, de-identified CT scans. We presented 627 challenging cases to 6 US-based and 6 Japan-based radiologists. In the experimental setup, readers were divided into two groups that read each case twice, with and without assistance from the model. Readers were asked to apply scoring guidelines they typically use in their clinical practice and report their overall suspicion of cancer for each case. We then compared the results of the reader’s responses to measure the impact of the model on their workflow and decisions. The score and suspicion level were judged against the actual cancer outcomes of the individuals to measure sensitivity, specificity, and area under the ROC curve (AUC) values. These were compared with and without assistance.
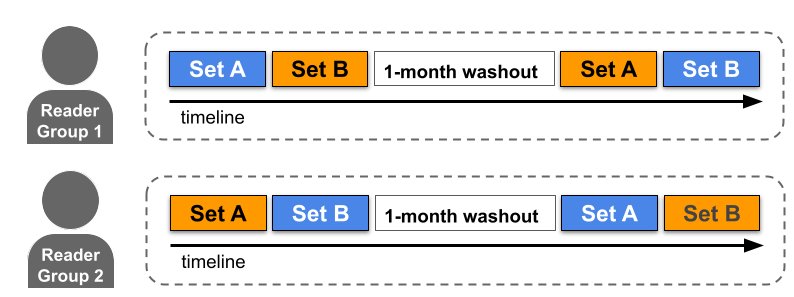
A multi-case multi-reader study involves each case being reviewed by each reader twice, once with ML system assistance and once without. In this visualization one reader first reviews Set A without assistance (blue) and then with assistance (orange) after a wash-out period. A second reader group follows the opposite path by reading the same set of cases Set A with assistance first. Readers are randomized to these groups to remove the effect of ordering.
The ability to conduct these studies using the same interface highlights its generalizability to completely different cancer scoring systems, and the generalization of the model and assistive capability to different patient populations. Our study results demonstrated that when radiologists used the system in their clinical evaluation, they had an increased ability to correctly identify lung images without actionable lung cancer findings (i.e., specificity) by an absolute 5–7% compared to when they didn’t use the assistive system. This potentially means that for every 15–20 patients screened, one may be able to avoid unnecessary follow-up procedures, thus reducing their anxiety and the burden on the health care system. This can, in turn, help improve the sustainability of lung cancer screening programs, particularly as more people become eligible for screening.
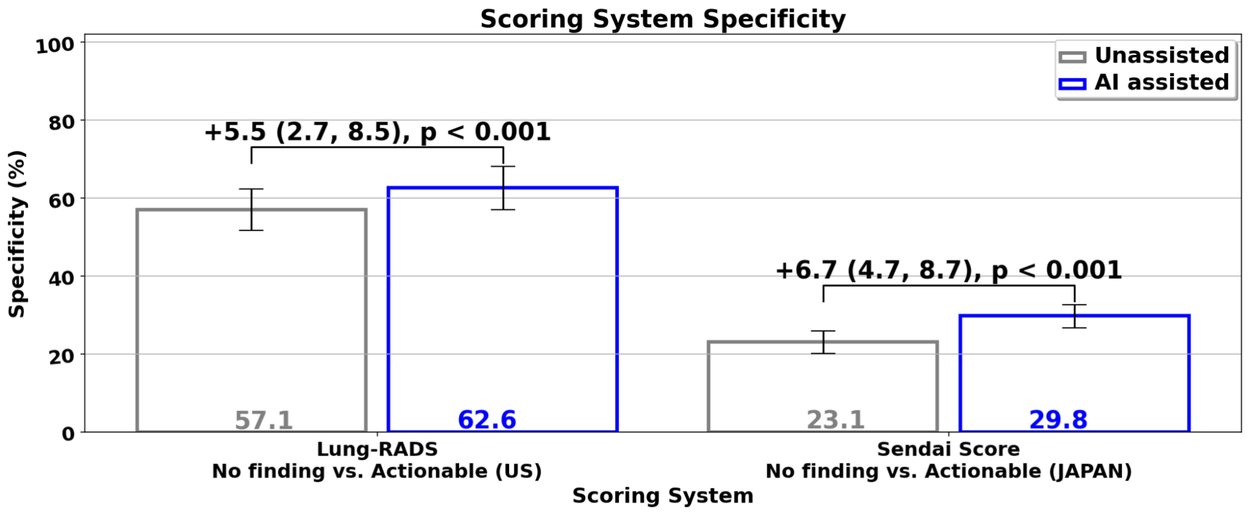
Reader specificity increases with ML model assistance in both the US-based and Japan-based reader studies. Specificity values were derived from reader scores from actionable findings (something suspicious was found) versus no actionable findings, compared against the true cancer outcome of the individual. Under model assistance, readers flagged fewer cancer-negative individuals for follow-up visits. Sensitivity for cancer positive individuals remained the same.
Translating this into real-world impact through partnership
The system results demonstrate the potential for fewer follow-up visits, reduced anxiety, as well lower overall costs for lung cancer screening. In an effort to translate this research into real-world clinical impact, we are working with: DeepHealth, a leading AI-powered health informatics provider; and Apollo Radiology International a leading provider of Radiology services in India to explore paths for incorporating this system into future products. In addition, we are looking to help other researchers studying how best to integrate ML model results into clinical workflows by open sourcing code used for the reader study and incorporating the insights described in this blog. We hope that this will help accelerate medical imaging researchers looking to conduct reader studies for their AI models, and catalyze translational research in the field.
Acknowledgements
Key contributors to this project include Corbin Cunningham, Zaid Nabulsi, Ryan Najafi, Jie Yang, Charles Lau, Joseph R. Ledsam, Wenxing Ye, Diego Ardila, Scott M. McKinney, Rory Pilgrim, Hiroaki Saito, Yasuteru Shimamura, Mozziyar Etemadi, Yun Liu, David Melnick, Sunny Jansen, Nadia Harhen, David P. Nadich, Mikhail Fomitchev, Ziyad Helali, Shabir Adeel, Greg S. Corrado, Lily Peng, Daniel Tse, Shravya Shetty, Shruthi Prabhakara, Neeral Beladia, and Krish Eswaran. Thanks to Arnav Agharwal and Andrew Sellergren for their open sourcing support and Vivek Natarajan and Michael D. Howell for their feedback. Sincere appreciation also goes to the radiologists who enabled this work with their image interpretation and annotation efforts throughout the study, and Jonny Wong and Carli Sampson for coordinating the reader studies.





