Announcing ScaNN: Efficient Vector Similarity Search
July 28, 2020
Posted by Philip Sun, Software Engineer, Google Research
Quick links
Suppose one wants to search through a large dataset of literary works using queries that require an exact match of title, author, or other easily machine-indexable criteria. Such a task would be well suited for a relational database using a language such as SQL. However, if one wants to support more abstract queries, such as “Civil War poem,” it is no longer possible to rely on naive similarity metrics such as the number of words in common between two phrases. For example, the query “science fiction” is more related to “future” than it is to “earth science” despite the former having zero, and the latter having one, word in common with the query.
Machine learning (ML) has greatly improved computers’ abilities to understand language semantics and therefore answer these abstract queries. Modern ML models can transform inputs such as text and images into embeddings, high dimensional vectors trained such that more similar inputs cluster closer together. For a given query, we can therefore compute its embedding, and find the literary works whose embeddings are closest to the query’s. In this manner, ML has transformed an abstract and previously difficult-to-specify task into a rigorous mathematical one. However, a computational challenge remains: for a given query embedding, how does one quickly find the nearest dataset embeddings? The set of embeddings is often too large for exhaustive search and its high dimensionality makes pruning difficult.
In our ICML 2020 paper, “Accelerating Large-Scale Inference with Anisotropic Vector Quantization,” we address this problem by focusing on how to compress the dataset vectors to enable fast approximate distance computations, and propose a new compression technique that significantly boosts accuracy compared to prior works. This technique is utilized in our recently open-sourced vector similarity search library (ScaNN), and enables us to outperform other vector similarity search libraries by a factor of two, as measured on ann-benchmarks.com.
The Importance of Vector Similarity Search
Embedding-based search is a technique that is effective at answering queries that rely on semantic understanding rather than simple indexable properties. In this technique, machine learning models are trained to map the queries and database items to a common vector embedding space, such that the distance between embeddings carries semantic meaning, i.e., similar items are closer together.
To answer a query with this approach, the system must first map the query to the embedding space. It then must find, among all database embeddings, the ones closest to the query; this is the nearest neighbor search problem. One of the most common ways to define the query-database embedding similarity is by their inner product; this type of nearest neighbor search is known as maximum inner-product search (MIPS).
Because the database size can easily be in the millions or even billions, MIPS is often the computational bottleneck to inference speed, and exhaustive search is impractical. This necessitates the use of approximate MIPS algorithms that exchange some accuracy for a significant speedup over brute-force search.
A New Quantization Approach for MIPS
Several state-of-the-art solutions for MIPS are based on compressing the database items so that an approximation of their inner product can be computed in a fraction of the time taken by brute-force. This compression is commonly done with learned quantization, where a codebook of vectors is trained from the database and is used to approximately represent the database elements.
Previous vector quantization schemes quantized database elements with the aim of minimizing the average distance between each vector x and its quantized form x̃. While this is a useful metric, optimizing for this is not equivalent to optimizing nearest-neighbor search accuracy. The key idea behind our paper is that encodings with higher average distance may actually result in superior MIPS accuracy.
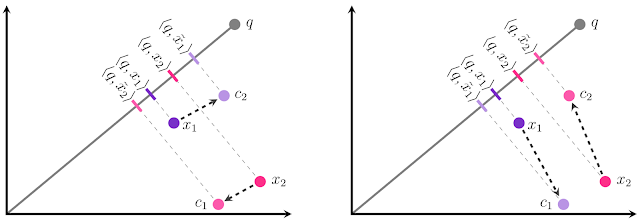
The intuition for our result is illustrated below. Suppose we have two database embeddings x1 and x2, and must quantize each to one of two centers: c1 or c2. Our goal is to quantize each xi to x̃i such that the inner product <q, x̃i> is as similar to the original inner product <q, xi> as possible. This can be visualized as making the magnitude of the projection of x̃i onto q as similar as possible to the projection of xi onto q. In the traditional approach to quantization (left), we would pick the closest center for each xi, which leads to an incorrect relative ranking of the two points: <q, x̃1> is greater than <q, x̃2>, even though <q, x1> is less than <q, x2>! If we instead assign x1 to c1 and x2 to c2, we get the correct ranking. This is illustrated in the figure below.
It turns out that direction matters as well as magnitude--even though c1 is farther from x1 than c2, c1 is offset from x1 in a direction almost entirely orthogonal to x1, while c2’s offset is parallel (for x2, the same situation applies but flipped). Error in the parallel direction is much more harmful in the MIPS problem because it disproportionately impacts high inner products, which by definition are the ones that MIPS is trying to estimate accurately.
Based on this intuition, we more heavily penalize quantization error that is parallel to the original vector. We refer to our novel quantization technique as anisotropic vector quantization due to the directional dependence of its loss function. The ability of this technique to trade increased quantization error of lower inner products in exchange for superior accuracy for high inner products is the key innovation and the source of its performance gains.
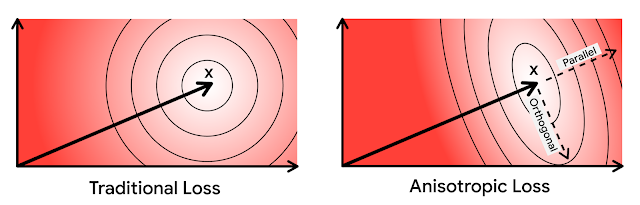 |
| In the above diagrams, ellipses denote contours of equal loss. In anisotropic vector quantization, error parallel to the original data point x is penalized more. |
Anisotropic Vector Quantization in ScaNN
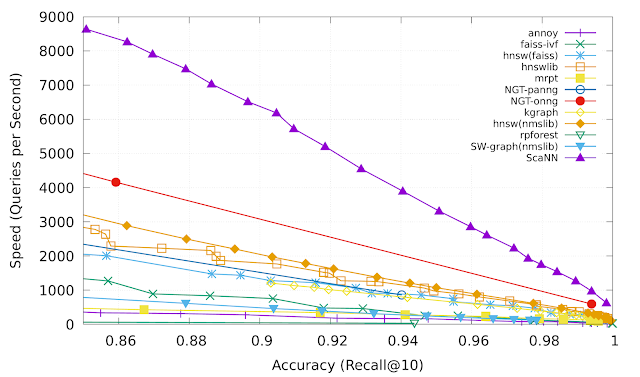
Anisotropic vector quantization allows ScaNN to better estimate inner products that are likely to be in the top-k MIPS results and therefore achieve higher accuracy. On the glove-100-angular benchmark from ann-benchmarks.com, ScaNN outperformed eleven other carefully tuned vector similarity search libraries, handling roughly twice as many queries per second for a given accuracy as the next-fastest library.*
ScaNN is open-source software and you can try it yourself at GitHub. The library can be directly installed via Pip and has interfaces for both TensorFlow and Numpy inputs. Please see the GitHub repository for further instructions on installing and configuring ScaNN.
Conclusion
By modifying the vector quantization objective to align with the goals of MIPS, we achieve state-of-the-art performance on nearest neighbor search benchmarks, a key indicator of embedding-based search performance. Although anisotropic vector quantization is an important technique, we believe it is just one example of the performance gains achievable by optimizing algorithms for the end goal of improving search accuracy rather than an intermediate goal such as compression distortion.
Acknowledgements
This post reflects the work of the entire ScaNN team: David Simcha, Erik Lindgren, Felix Chern, Nathan Cordeiro, Ruiqi Guo, Sanjiv Kumar, and Zonglin Li. We’d also like to thank Dan Holtmann-Rice, Dave Dopson, and Felix Yu.
* ScaNN performs similarly well on the other datasets of ann-benchmarks.com, but the website currently shows outdated, lower numbers. See this pull request for more representative performance figures on other datasets. ↩